TCRMatch - Tutorial
Note on large datasets
Currently, the IEDB-AR is not equipped with hardware to accomodate input datasets of >500 CDR3b sequences as input for TCRMatch. However, two options are available
for analyzing large datasets:
1. TCRMatch is available at VDJServer.org following the creation of a free account.
2. TCRMatch can be run locally as a command line tool. Please click here to download the stanalone, and visit our github respository for more information.
1. TCRMatch is available at VDJServer.org following the creation of a free account.
2. TCRMatch can be run locally as a command line tool. Please click here to download the stanalone, and visit our github respository for more information.
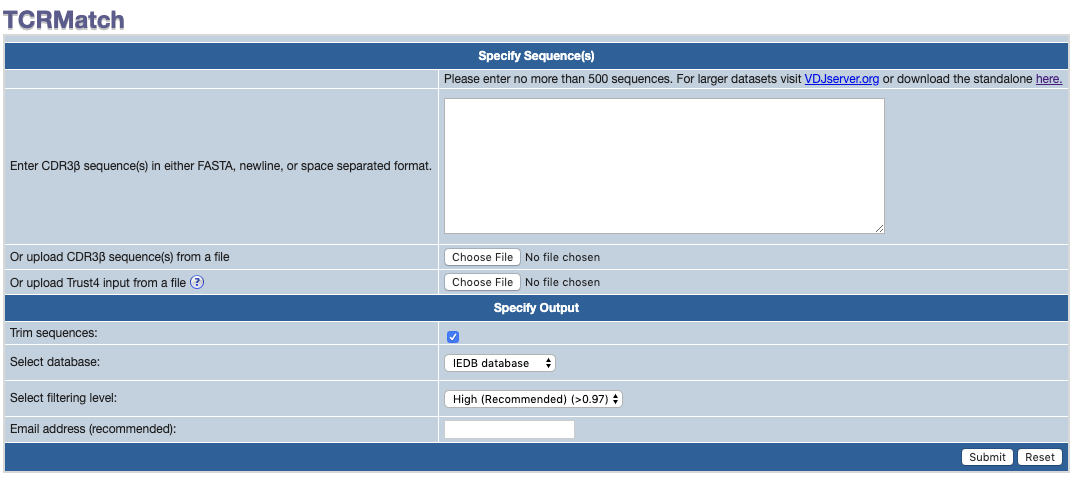
1. Specify sequences to query against the IEDB or CEDAR database
First specify the sequences you want to query against the database.
The sequences should be entered directly into the text area, or can be taken from a file that is uploaded through the button.
Please enter no more than 500 sequences at a time. If you need to perform large queries, we recommended downloading the standalone version and running locally.
The sequences can be provided as either FASTA format, or new line separated sequences.
• Upload Trust4 file instead of specifying sequences:
Users can upload a TRUST4 file as input instead of Specifying sequences. TCRMatch filters out IG genes from the TRUST4 file before running the comparisons. Tcr Receptor Utilities for Solid Tissue (TRUST) is a computational tool to analyze TCR and BCR sequences using unselected RNA sequencing data, profiled from solid tissues, including tumors. TRUST4 performs de novo assembly on V, J, C genes including the hypervariable complementarity-determining region 3 (CDR3) and reports consensus of BCR/TCR sequences. TRUST4 then realigns the contigs to IMGT reference gene sequences to report the corresponding information. TRUST4 supports both single-end and paired-end sequencing data with any read length.
2. Select optional sequence trimming
This parameter removes the conserved C/F+W motif from input sequences if selected.
3. Select Database
Select the database (IEDB or CEDAR) to query against.
4. Select stringency level
TCRMatch operates by generating a score between 0 and 1, with 1 being an exact match to the input sequence. The score selected will be used to filter out any sequences
that score less than this number. We recommend using a threshold of .97, which provides a good balance of precision and recall. Lower values are provided, but may
result in returning unrelated CDR3b sequences. The selection "show all peptides" returns all sequences from the IEDB and their corresponding TCRMatch scores.
5. Provide an email address (optional)
Providing an email in this field allows us to send the results of the search through email as a CSV file attachment.
This is recommended if you experience connectivity issues, as you can navigate away from the page and still receive results. This is also ideal for large input
datasets which may take time to run. An email is required for all datasets larger than 100 sequences.
